A Machine Learning approach to the precision treatment of chronic osteoarthritis pain
Overview
InSyBio is a biotechnology company that provides precise solutions for biomarker discovery in the nutrition industry, biotechnology industry, and academia. It has developed a machine-learning tool for the precision treatment of chronic pain using non-opioid treatments for patients suffering from osteoarthritis, a degenerative condition and one of the primary causes of chronic pain worldwide. Nikos Iliopoulos and Argiris Karageorgos worked on the project and created prediction models that will be integrated into a clinical decision support system and allow for the personalization of osteoarthritis treatment. Within the framework of this project, they created:
- Models for the precision treatment of osteoarthritis patients.
- Prediction scripts that will be implemented in InSyBio's software and facilitate the personalization of osteoarthritis treatment.
Case Study Contents
- The background
- The goal of the project
- Tools & Methodology
- Getting started
- Conclusions & Recommendations
- Online presentation
1. The background
InSyBio is a bioinformatics company meeting the needs of molecular biology researchers. It focuses on developing computational frameworks and tools for the analysis of complex life-science and biological data.
The company aims to accelerate preclinical and clinical studies of pharmaceutical companies and provide precise solutions for biomarker discovery in the nutrition industry, biotechnology industry, and academia.
InSybio offers tools for
- biomarker discovery
- biological data management
- secured storage
- computational bio-data analysis.
InSybio is striving for the continuous production of intelligent and innovative algorithms that assist in inferring knowledge from experimental data. To that end, the company collaborated with Big Blue Data Academy, and Nikos Iliopoulos and Argiris Karageorgos worked on A precision medicine approach for treating osteoarthritis for their final project.
2. The goal of the project
The work of Nikos and Argiris aims to utilize InSyBio's machine learning methodology and other machine learning techniques combining:
- multi-objective optimization algorithms and
- classification/regression models (gradient boosting, neural networks, random forests, etc.
and train models that can predict the outcome of osteoarthritis patients considering their:
- clinical features
- responses to questionnaires
- medication.
These models will then be integrated into a precision treatment system to identify the best performing treatment for osteoarthritis patients. The milestones of the project were:
Milestone 1: Develop and evaluate osteoarthritis treatment outcome prediction models for a given set of medications using machine learning.
Milestone 2: Integrate these prediction models into a clinical decision support system that will allow for the personalization of osteoarthritis treatment.
3. Tools & Methodology
In addition to the proprietary machine-learning tool, InSybio has data from the Osteoarthritis Initiative Database and other population observational pain studies. This data includes clinical, imaging, and questionnaire data for more than 5000 patients and their 6-1 year follow-up regarding the evaluation of the pain condition and whether surgery was required or not, considering that different medications were used. The tools and methodology of the project included:
- Machine learning python libraries - random forests, gradient boosting, and deep learning methods
- Software of InSyBio - ML implementations and InSyBio Biomarkers tool for data preprocessing
Methodology
The methodology included:
- Exploratory Data Analysis
- Feature Engineering
- ML Algorithms (Classification, Regression)
- Resulting scores
4. Getting started
The team worked on two datasets that were based on the patients' answers to osteoarthritis questionnaires.
- Prevent baseline dataset
- Clinical review dataset
Our pipeline - Prevent baseline dataset - Argiris Karageorgos
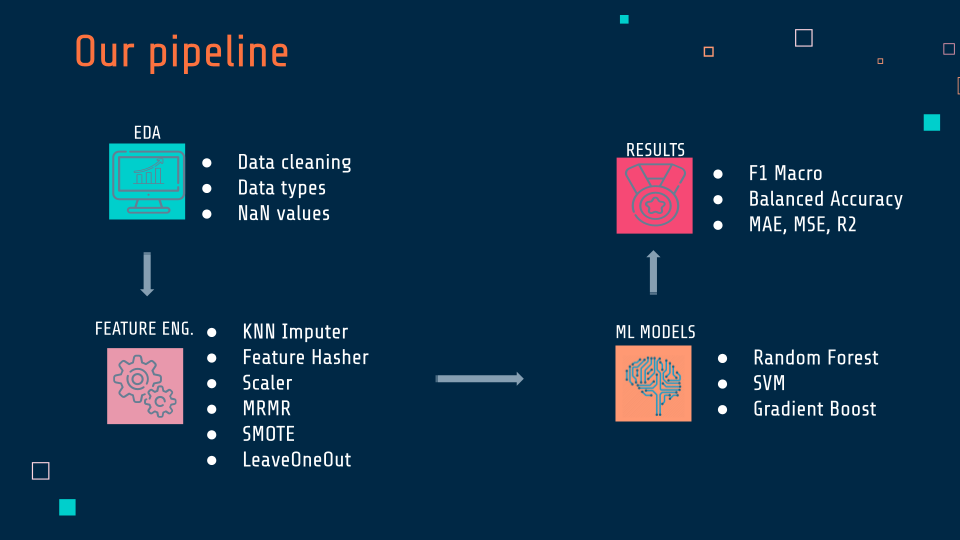
The first step was to clean the data.
In some columns, values had to be changed from numerical to categorical, so there was a better match to the patients' answers. Then some columns were dropped, which had a high percentage of sine values, and for the NaN that remained, the values were filled in using KNN Imputer. Using the Feature Hasher, the team was able to turn the categoricals into numericals and work on them.
Scaling was done on the dataset, and through MRMR, they got the best features for their models. Since the datasets were unbalanced, the team used SMOTE, which they applied to the grid search they created and did cross-validation within the grid-search with the LeaveOneOut technique. They applied this grid search to the Random Forest model, SVMand Gradient Boost, where they got the optimal parameters and the best scores through the metrics of F1 Macro, Balanced Accuracy, MAE, MSE, and R2.
Clinical review dataset – Nikos Iliopoulos
.png)
In the clinical review dataset, there were both regressors and classifiers. Specifically, the classifier, as a target in the column, if the patient had surgery after five years. For that target, the team used the three algorithms SVM, Gradient Boost, Random Forest. A grid search was done to find the optimal parameters in all algorithms, and the score was done based on F1 Macro.
- Gradient Boost was in first place with a very good score on F1 Macro with 0.95.
- Balanced Accuracy also 0, 95.
Once the first task was completed for the classifier, the team looked into what impact a column has on whether patients had surgery after five years. SHAP values were implemented.
The first place was given to pain, whether - patients have terrible pain in their daily lives. Then, age plays a role, as well as whether patients do any physical exercise.
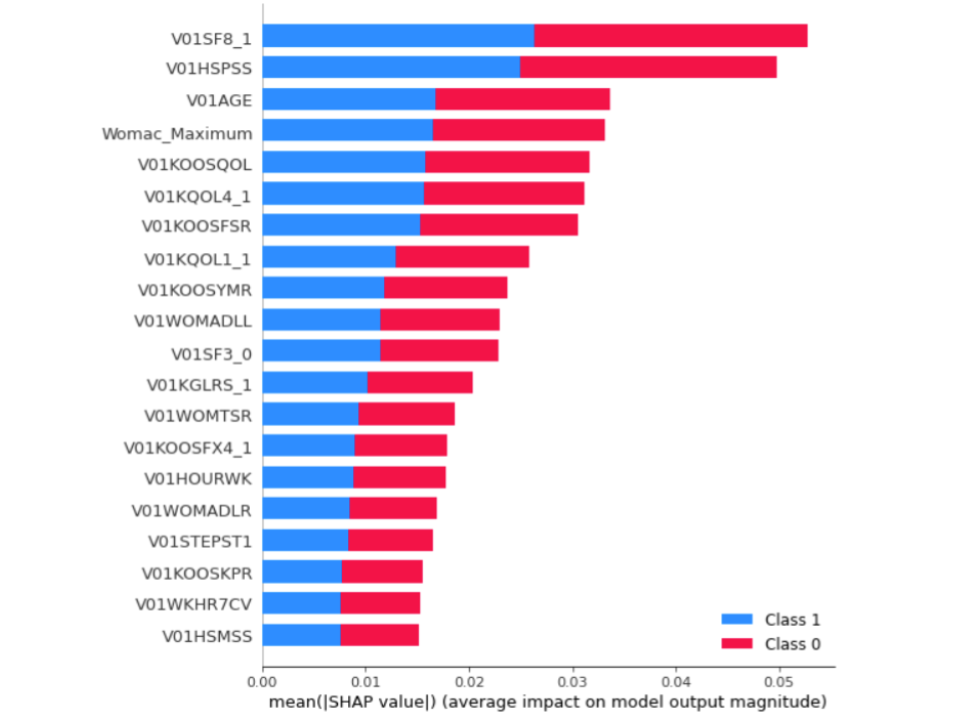
SHAP Values
V01SF8_1: How much did pain interfere with normal work
V01HSPSS: Physical summary scale for the MOS 12-item short-form health survey
V01AGE: Age
V01KOOSQOL: KOOS Quality of Life Score
V01KOOSFSR: Sports and Recreational Activities Score
.1.png)
There were two regressors, two scores:
- quality of life
- pain score
The team presented the maximum pain score because the results were similar in both regressors. They applied the same algorithms with the difference with linear regression. Again, a grid search was done to find the optimal parameters describing our model. The score was based on MSE because we had several outliers. Random Forest took first place with a very good score in the third decimal place, 0.003, MAE was at 0.04, and R2 was at 0.19.
As with the classifiers, the team looked into the columns that had the most significant impact on the target group. Since the target group was about pain, the biggest impact is how much pain the patient feels in their daily lives. What we could look at next is again the physical exercise of each patient and, interestingly, body weight, Body Mass Index, now plays a role.
.png)
Most important features
Womac_Maximum: How much did pain interfere with normal work
V01HSPSS: Physical summary scale for the MOS 12-item short-form health survey (SF-12)
V01KOOSFSR: Sports and Recreational Activities Score
V01BMI: Body mass index
V01PASE: Physical Activity Scale for the Elderly score
Prevent Baseline Database
In the prevent baseline dataset, there were three columns for output: severity decrease, interference decrease, and TotalMeds decrease. They were all binary, so the team used the three algorithms mentioned above, Random Forest, Gradient Boost, and SVM. All three were applied to all outputs, and the best score was Random Forest with 0.88, for the second output, Gradient Boost 0.90, for the third, SVM 0.88.
.%5B1%5D.png)
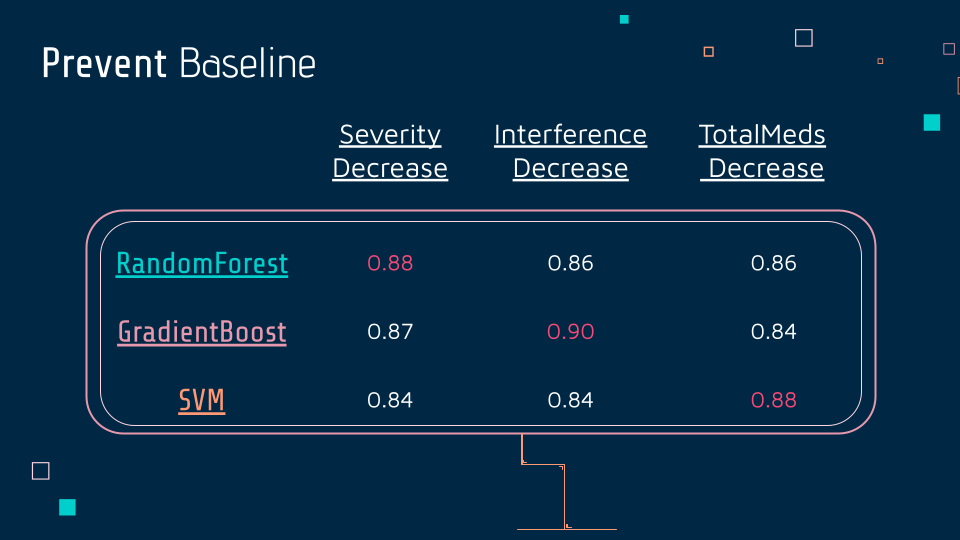
Summary of Qualitative Results
| Target: Severity Decrease |
Target: Interference Decrease |
Target: TotalMeds Decrease |
|
Random Forest: 0.88 |
Random Forest: 0.86 |
Random Forest: 0.86 |
|
Gradient Boost: 0.87 |
Gradient Boost: 0.90 |
Gradient Boost: 0.84 |
| SVM: 0.84 | SVM: 0.84 | SVM: 0.88 |
| Applied ML models for classification |
Applied ML models for regression |
Extracted most important features for classifiers and regressors |
|
Best estimator: Gradient Boost |
Best estimator: Random Forest |
|
| F1 macro: 0.95 | MSE: 0.003 | |
| Accuracy: 0.95 | MAE: 0.04 | |
| Balanced Accuracy: 0.95 |
Prevent Baseline analysis
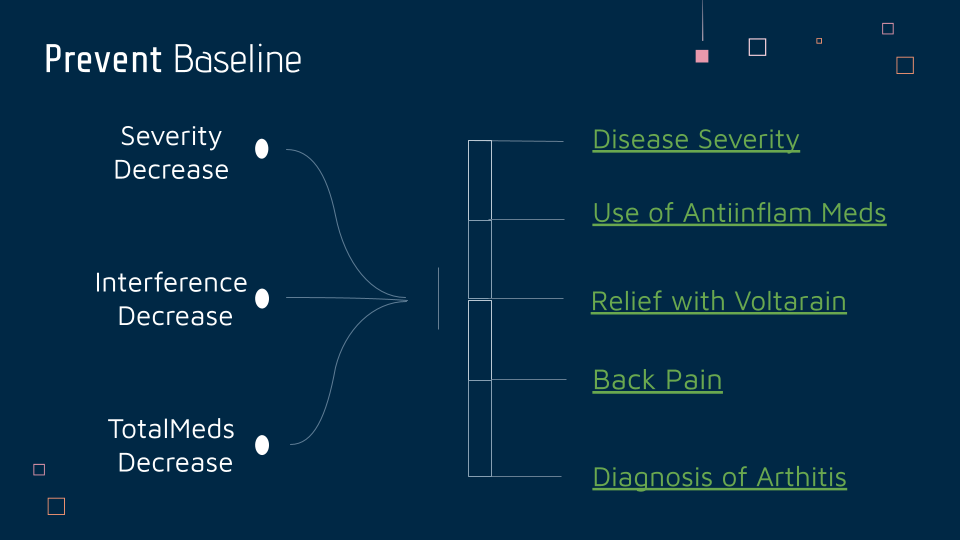
Then, the team used several features to predict all three targets, but some were common to all three and were very important. The common features were whether:
- the patient's daily life was affected by the problem
- patients were using anti-inflammatory drugs
- patients were using Voltaren
- the pain was in the back of the body
- the diagnosis for arthritis was positive
5. Conclusions & Recommendations
After the pipeline with the preprocess was built, the team took the best models, and based on them, they created prediction scripts for InSyBio's software. The prediction models will be integrated into a clinical decision support system that will allow for the personalization of osteoarthritis treatment. Medical experts will test the treatments at the Patras University Hospital.
For the continuation of the project and its further improvement, the team recommends using the algorithms of CatBoost, XGBoost, and Classifier Chain. The team used them and saw great potential for further improvement since the tests provided excellent results.
6. Online presentation
You can see the final data science project presentation here.
For more information on Big Blue Data Academy projects, you can contact us here.
Invitation
Companies in Greece are invited to participate in the Big Blue Training Process and assign data-driven projects to Bootcamp students.


.jpg)